SOMAmer (Slow Off‑rate Modified Aptamer) technology is an advanced proteomics tool that leverages next‑generation sequencing (NGS) to enable highly multiplexed protein measurement. SOMAmer Reagents are chemically modified, single-stranded DNA molecules developed for exceptional specificity and affinity for a target protein through precise three-dimensional shape complementarity. These structures accurately bind protein targets with high sensitivity, allowing researchers to quantify thousands of proteins simultaneously in a single assay. Because SOMAmer Reagents are designed rather than biologically produced, they offer more consistent performance, higher reproducibility, and improved batch‑to‑batch reliability compared to traditional antibody-based proteomics methods.1
What is SOMAmer technology?

What are the benefits of using SOMAmer technology for protein detection?
SOMAmer-based proteomics assays deliver a powerful combination of specificity, scalability, and reproducibility, enabling researchers to accurately explore the proteome.
Key advantages of SOMAmer technology are:
- Scalability: Scale high‑plex SOMAmer‑based assays to thousands of protein targets with consistent performance and minimal lot‑to‑lot variability enabled by synthetic, sequence‑defined reagents
- Exceptional precision: Enhance the power of high-content assays to reveal rare biomarkers and subtle disease-associated signals with dependable results
- High sensitivity: Enable quantification of proteins across an exceptionally broad range down to femtomolar detection with > 10-log dynamic range2-5
- Outstanding reproducibility: Obtain reliable and biologically meaningful insights using fully synthetic reagents with consistently low CVs (~5.5%)
Advancing high-throughput proteomics with NGS
Download this eBook to learn how high-throughput proteomics is now more accessible than ever with the Illumina Protein Prep solution.
Download eBook
Case studies and applications
Proteomics and rare diseases
Dr Matthew Brown, Chief Scientific Officer at Genomics England, shares results from a pilot project with Illumina Protein Prep, screening ~1500 participants with rare diseases to demonstrate the power of NGS-based proteomics for biomarker discovery research.
This poster showcases the Illumina Protein Prep assay’s exceptional sensitivity and its strong concordance with SomaScan data from SomaLogic. The ability to detect abundant differential proteins across specific disease categories demonstrates how high‑resolution proteomics can deepen our understanding of rare genetic conditions by revealing disease‑associated genes and molecular pathways.
Rare melanoma research powered by Illumina Protein Prep
Watch how Assistant Professor Kacey Couts and her team at the University of Colorado are using Illumina Protein Prep to uncover novel insights into rare melanomas. This assay identifies 9.5K unique human proteins across more than 200 biological pathways, enabling her team to maximize discovery power as they search for novel biomarkers and investigate treatment responses.
In this podcast, the Associate Director for Product Management at Illumina talks about SomaLogic, the launch of Illumina Protein Prep, and its link to the broader multiomics goal to improve biomarker discoveries and phenotypic insights.
SOMAmer Reagents vs antibodies: Why researchers are turning to SOMAmer Reagents for affinity-based proteomics
SOMAmer Reagents are fully synthetic oligonucleotides that bind proteins with high affinity and specificity, whereas antibodies are biologically derived proteins that have inherent variability. The synthetic design of SOMAmer Reagents enables superior reproducibility, enhanced detection sensitivity, and ultra‑high multiplexing, making them well‑suited for the demands of modern high‑throughput proteomic applications.
SOMAmer Reagent
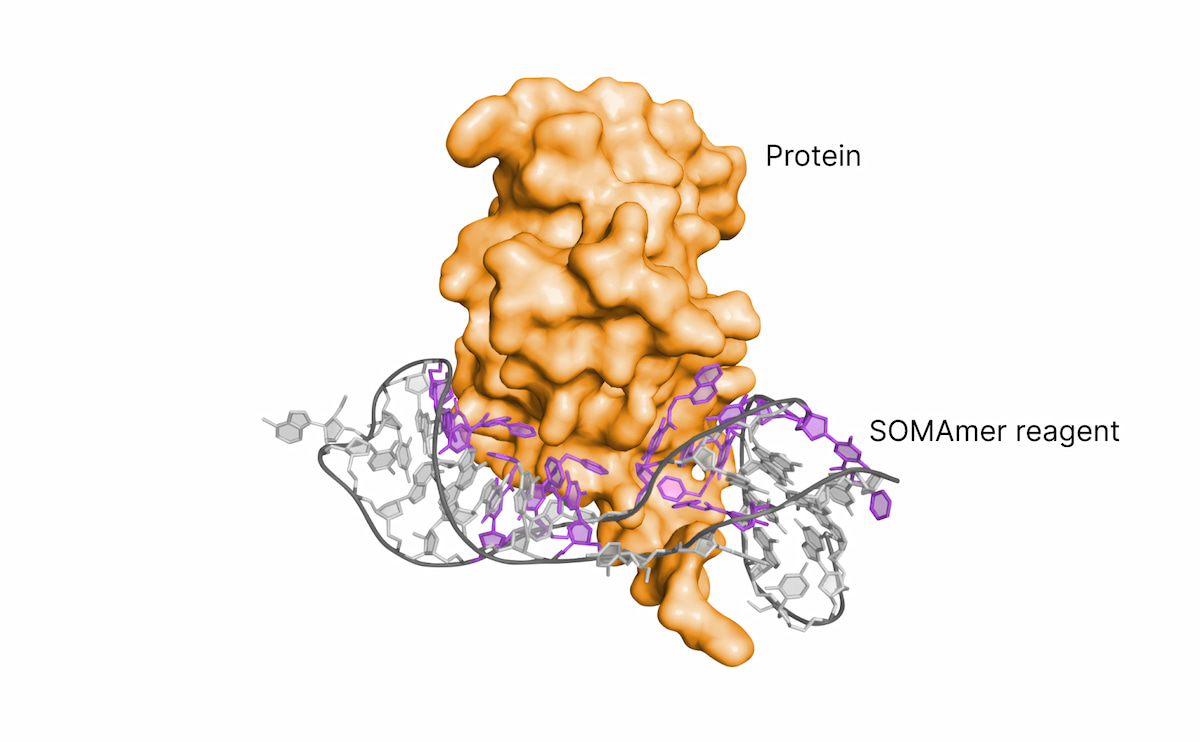
- Short, single-stranded DNA with modified nucleotides (~20 kDa)
- Chemically synthesized with a consistent structure
- Multiplexing without limitations
- Polyanionic competitors are used to block transient interaction reducing cross reactivity with non-target proteins
- Content scalability
Antibody
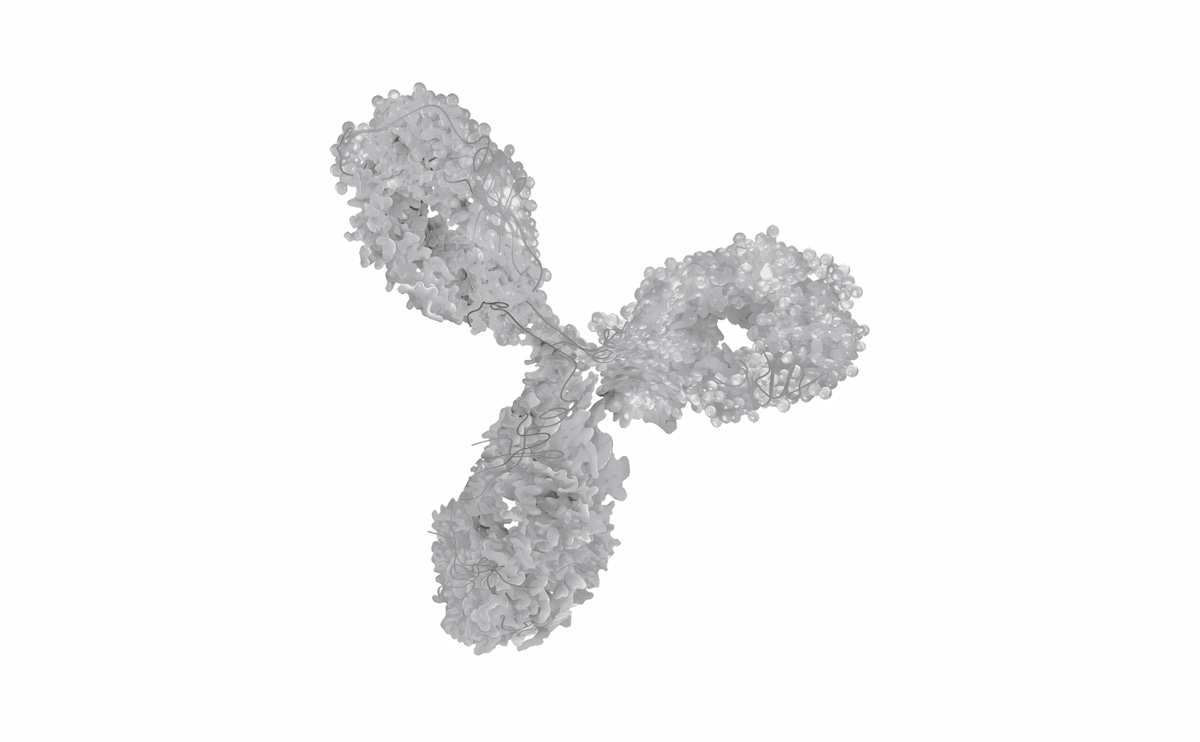
- Large, complex protein (~150 kDa)
- Produced biologically with natural structural variability
- Limited multiplexing options
- No universal solution to reduce non-specific binding, leading to higher cross-reactivity with non-target proteins
- Limited scalability

Precision proteomics with Illumina Protein Prep
How do SOMAmers power NGS-based proteomics?
SOMAmer Reagents enable high-precision and high-throughput proteomics by linking protein detection with the proven scalability of next-generation sequencing (NGS). Each SOMAmer Reagent binds a specific protein and carries a unique DNA tag. After capture, these tags are converted into sequencing libraries, allowing protein abundance to be quantified by counting sequencing reads using an NGS platform. This approach transforms proteomics into a high-throughput automated workflow, making it possible to profile thousands of proteins in parallel with the same efficiency and precision researchers expect from other NGS-based methods.
Achieve deeper biological insights with integrated proteomics and NGS
SOMAmer-based proteomics is now integrated into Illumina NGS sequencing workflows, combining the value of proteomic analysis with other omic data. This synergy allows scientists to connect protein-level insights with genomic and transcriptomic data, providing a more complete picture of biological systems and accelerating discoveries that can inform precision medicine, biomarker discoveries, drug targets, disease pathways, large-scale studies, and more with high-throughput scalability.
Illumina Protein Prep workflow
Using Illumina Protein Prep, SOMAmer Reagents capture thousands of proteins in a single reaction. These protein-binding events are converted into sequencing-ready libraries through a streamlined, automated process.
Prep
This automated NGS-based proteomics solution with SOMAmer technology and Illumina sequencing and analysis detects and quantifies 9.5K human proteins in 2.5 days.
Sequence
The NovaSeq X Series delivers extraordinary throughput and accuracy to perform data-intensive applications at production scale.
NovaSeq 6000 System expands sequencing capabilities by combining throughput, flexibility, and simplicity for virtually any method, genome, and scale.
Data analysis
Integrated secondary analysis pipeline auto-launches after sequencing for complete normalization and protein expression quantification.
Interpretation
Illumina Connected Multiomics is a powerful and intuitive cloud-based solution that enables multiomic data analysis and visualization at scale.
Access NGS-based proteomics through Illumina Protein Prep services
You can now take advantage of Illumina Protein Prep without installing a full workflow in-house. Through a network of core labs and trusted service providers partnering with Illumina, you can outsource high-throughput proteomics projects and receive accurate, reproducible results powered by SOMAmer technology and Illumina sequencing. This service option removes barriers to adoption, enabling labs of any size to explore the proteome at scale whether for biomarker discovery, drug development, or multiomic studies, without the need for specialized infrastructure.

FAQ
Traditional antibodies are large affinity reagents (~150 kDa) that typically bind three‑dimensional structures on the surface of proteins that have triggered an immune response (ie, immunogenic targets). They are produced biologically either as monoclonal antibodies generated in cell culture to recognize a single protein epitope, or as polyclonal antibodies produced in animals that recognize multiple epitopes on the same protein.
SOMAmer Reagents, in contrast, are small, chemically modified aptamers (8—20 kDa) that also bind to three‑dimensional protein structures. Because they are fully synthetic, they provide consistent batch‑to‑batch performance. Their distinct chemistry also allows them to bind non‑immunogenic and epitopes currently not targeted by antibodies.
Together, these features make SOMAmer Reagents a versatile and reliable alternative to traditional antibody‑based tools.
SOMAmer‑based proteomics enables high‑throughput, cost‑effective workflows with > 10‑log dynamic range, femtomolar sensitivity, and scalable multiplexing, all with lower variability than traditional methods. This allows reliable detection of low‑abundance proteins without requiring any changes to the workflow and makes SOMAmer‑based proteomics a powerful and efficient approach for broad, reliable protein measurement at scale.
In contrast, mass spectrometry typically requires additional sample processing steps to improve coverage of lower‑abundance proteins.
Compared to antibody‑based approaches, SOMAmer‑based proteomics consistently delivers strong reproducibility (~5% CVs), even with expanded assay multiplexing.
The workflow takes about 2.5 days from sample to analysis, with less than 4 hours of hands-on time.
Each SOMAmer Reagent achieves its specificity through two key properties: high‑affinity binding to its target protein and a slow dissociation rate. These characteristics stem from the reagent’s unique chemical structure.
Before a SOMAmer Reagent is used in an assay, it undergoes primary validation, which includes pull‑down experiments and dose‑response assays to confirm that it binds selectively to its intended target. In addition, more than 70% of SOMAmer Reagents have supporting specificity data generated by independent researchers using orthogonal validation techniques, such as mass spectrometry.
SOMAmer Reagents are synthetic and can be consistently regenerated, ensuring lot-to-lot consistency and higher reproducibility than other affinity based methods.
Speak with a specialist
Interested in learning more about how to collaborate with Illumina or want to know more about instrument insourcing solutions? Fill out the form and a specialist will be in touch with you soon.
Speak to a specialist
References
- Brody EN, Gold L. Aptamers as therapeutic and diagnostic agents. J Biotechnol. 2000;74(1):5-13. doi:10.1016/s1389-0352(99)00004-5
- Eldjarn GH, Ferkingstad E, Lund SH, et al. Large-scale plasma proteomics comparisons through genetics and disease associations. Nature. 2023;622(7982):348-358. doi:10.1038/ s41586-023-06563-x
- Rooney MR, Chen J, Ballantyne CM, et al. Plasma proteomic comparisons change as coverage expands for SomaLogic and Olink. Preprint. medRxiv. 2024;2024.07.11.24310161. doi:10.1101/2024.07.11.24310161
- Gold L, Ayers D, Bertino J, et al. Aptamer-based multiplexed proteomic technology for biomarker discovery. PLoS One. 2010;5(12):e15004. doi:10.1371/journal.pone.0015004