Compare whole-genome and exome sequencing
Explore the benefits of each approach to determine which method is best for your research.
Whole-genome sequencing (WGS) is a comprehensive method for analyzing entire genomes. Genomic information has been instrumental in identifying inherited disorders, characterizing the mutations that drive cancer progression, and tracking disease outbreaks. Rapidly dropping sequencing costs and the ability to produce large volumes of data with today’s sequencers make whole-genome sequencing a powerful tool for genomics research.
While this method is commonly associated with sequencing human genomes, the scalable, flexible nature of next-generation sequencing (NGS) technology makes it equally useful for sequencing any species, such as agriculturally important livestock, plants, or disease-related microbes.
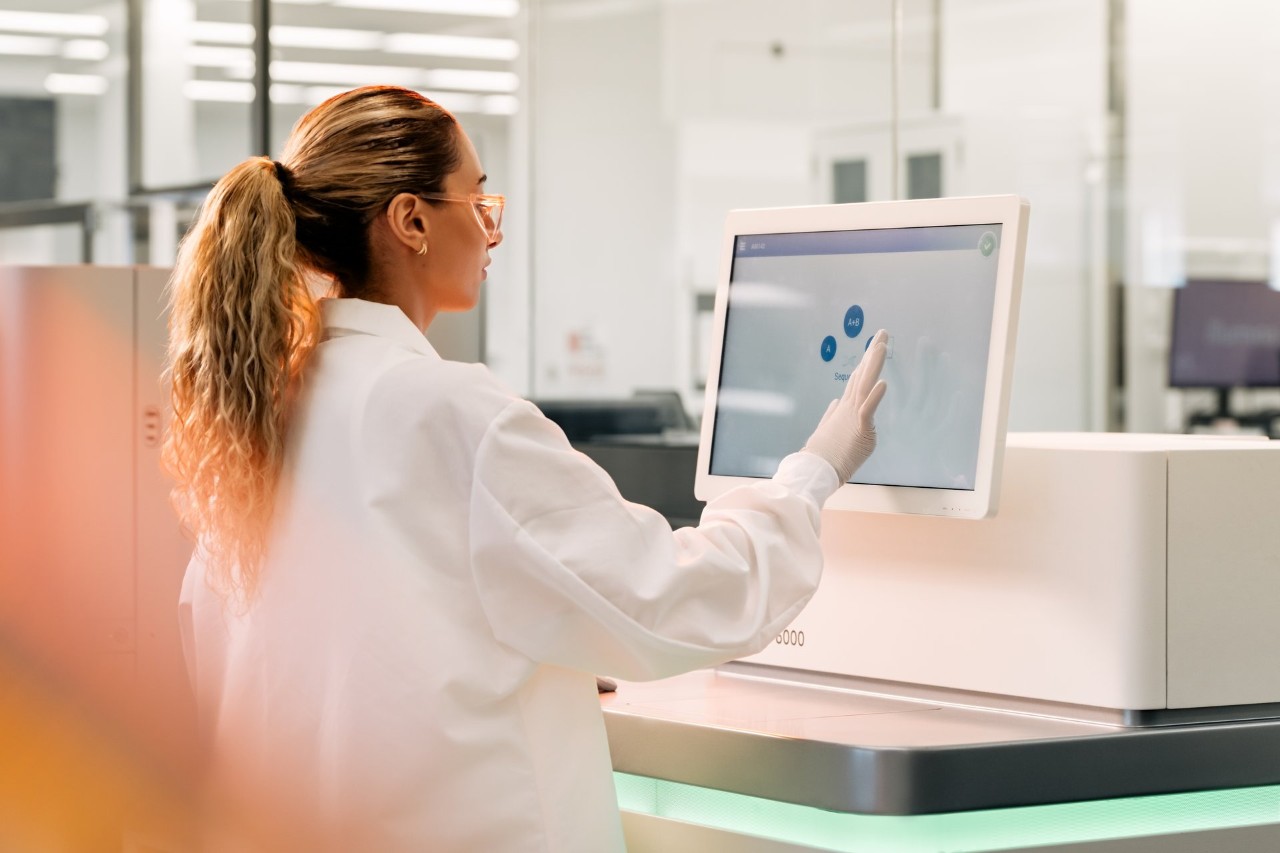

Unlike focused approaches such as exome sequencing or targeted resequencing, which analyze a limited portion of the genome, whole-genome sequencing delivers a comprehensive view of the entire genome. It is ideal for discovery applications, including identifying causative variants and novel genome assembly.
Whole-genome sequencing can detect single nucleotide variants, insertions/deletions, copy number changes, and large structural variants. Due to recent technological innovations, the latest genome sequencers can perform whole-genome sequencing more efficiently than ever.
Compare whole-genome and exome sequencing
Explore the benefits of each approach to determine which method is best for your research.
Sequencing large genomes (> 5 Mb), such as plant, animal, or human genomes, can provide valuable information for disease research and population genetics.
Small genome sequencing (≤ 5 Mb) involves sequencing the entire genome of a bacterium, virus, or other microbe. Without requiring bacterial culture, researchers can sequence thousands of small organisms in parallel using NGS.
De novo sequencing refers to sequencing a novel genome where there is no reference sequence available. NGS enables fast, accurate characterization of any species.
Phased sequencing, or genome phasing, distinguishes between alleles on homologous chromosomes to resolve their inheritance, resulting in whole-genome haplotypes. This information is often important for genetic disease studies.
Previously a challenging application, human whole-genome sequencing has never been simpler. It offers the most detailed view into our genetic code.
Long reads can help resolve challenging regions of the genome, such as those containing highly variable or highly repetitive elements.
Learn how Illumina innovations are redefining the limits of what's possible in genome sequencing. Dr Steve Barnard, Dr Joe Devaney, and Dr Bekim Sadikovic discuss groundbreaking technologies that simplify workflows and expand the frontiers of discovery.
This user-friendly three-step WGS workflow provides a fully featured, rapid solution for labs and delivers high-quality insights across the entire genome for all variant classes.
A fast, integrated workflow for a wide range of applications, from human whole-genome sequencing to amplicons, plasmids, and microbial species.
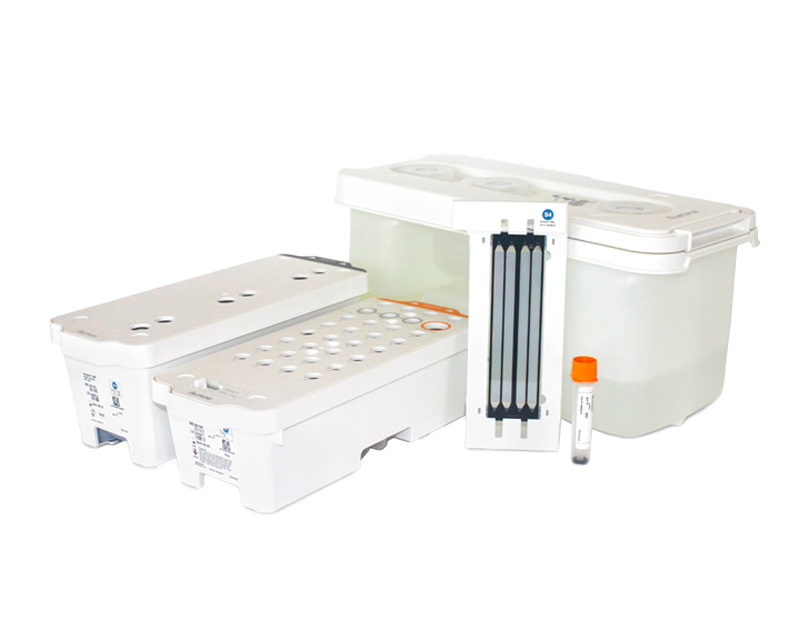

Reagent kits for the NovaSeq 6000 System provide ready-to-use cartridge-based reagents for cluster generation and SBS.

Data management and simplified bioinformatics for labs getting started and for rapidly scaling next-generation sequencing operations.

See how you can use multiomics to better connect genotype to phenotype and obtain a full cellular readout not found through single omic approaches.
WGS for Parkinson's disease research
Learn how NysnoBio scientists use whole-genome sequencing to study early-onset Parkinson’s disease.
The time is now for microbiome studies
Whole-genome shotgun sequencing and transcriptomics provide researchers and pharmaceutical companies with data to refine drug discovery and development.
Analyzing leukemia samples with WGS
A New England Journal of Medicine study found that using WGS to assess leukemia samples produced more accurate results, in less time, than karyotyping or fluorescence in situ hybridization.
Unlock long-distance genomic information, resolve challenging-to-map regions, and simplify your sequencing workflow with proximity mapped read technology.
Benefits of WGS for drug discovery and complex disease research
Nature study demonstrates the advantages of Illumina WGS over whole-exome sequencing and genotyping arrays for assessing genetic risk and identifying drug target candidates.
From samples to genomic insights
Learn about Illumina solutions for connecting samples to genomic analysis and interpretation in this recorded webinar.
WGS is a method that is used to gain comprehensive insights by analyzing entire genomes. Advancements in next-generation sequencing coupled with the flexible and scalable nature of NGS technologies make WGS useful for studying genetic material from humans, animals, plants, microbes, and viruses.1
Overview of an NGS workflow:
Whole-genome sequencing plays a critical role in human health. WGS helps researchers understand mutations in tumors that may drive cancer, assess inherited disease risk and carrier status, and gain insights into the genetic makeup of individuals for personalized medicine studies.
In public health, WGS is used to conduct infectious disease research, detect and track disease outbreaks, monitor the spread of infectious diseases, and identify emerging variants. WGS is also used in basic genetics research.1
Watch our WGS webinar and learn about accelerating precision medicine research with low-cost whole-genome sequencing technologies.
Sequencing approaches range from targeted to comprehensive. Targeted sequencing focuses on specific genes or genomic regions, typically selected based on a defined indication. Whole-exome sequencing broadens this scope by assessing multiple variant types across the protein coding regions of the genome. In contrast, whole-genome sequencing provides a comprehensive view of the entire genome, enabling detection of single nucleotide variants (SNVs), insertions and deletions, copy number variations, and large structural variants.
Some limitations of WGS include technical difficulties with data handling and interpretation of certain regions of the genome such as noncoding regions. There are also challenges with the use of some short-read technology for WGS and its ability to resolve long repetitive regions or structural variants.2
While technically a short-read technology, proximity mapped read technology, suitable for WGS needs, generates accurate long-range genomic insights.
The unique workflow maintains the link between the original long DNA template and the resulting short sequencing reads, enabling enhanced detection of structural variants, ultra-long phasing of genetic variants, and improved mapping in low-complexity regions.
Learn more about proximity mapped read technology and our DRAGEN secondary analysis pipeline.
To help you meet your research objectives, we offer a range of user-friendly primary, secondary, tertiary, and cloud-based sequencing data analysis solutions. For additional information, visit our sequencing data analysis resource hub or the DRAGEN secondary analysis page to find the right solution for your data needs.
Cancer whole-genome sequencing
Get a base-by-base view of cancer-associated variants.
Rare disease whole-genome sequencing
Learn how WGS can uncover rare disease variants.
Our innovative next-generation sequencing (NGS) platforms deliver exceptional data quality and accuracy at a massive scale. View benchtop and production-scale sequencers and find resources designed to help you choose the right platform.
Accurate CYP2D6 genotyping using WGS data
WGS offers a promising option for building accurate drug-metabolizing enzyme allele frequency databases for pharmacogenomics.
Library Prep and Array Kit Selector
Determine the best kit for your needs based on your project type, starting material, and method of interest.
Find high-quality whole-genome and other sequencing services that deliver analyzed data to researchers.
Use this interactive tool to explore experimental NGS library preparation methods compiled from the scientific literature.
Learn how to estimate and achieve the necessary sequencing coverage for your experiment.
Talk to an expert to learn more about solutions for whole-genome sequencing.